Introduction
Degenerative disorders of the spinal cord are significant health issues faced by the global population, and most cases of degenerative disorders of the spinal cord are triggered by intervertebral disc degeneration (IDD) [1, 2]. While the detailed mechanisms underlying the pathogenesis of IDD remain unclear, major progress has been made to elucidate relevant molecular pathways involved in the pathogenesis of IDD. The onset of IDD is a process involving multiple factors, including a critical step involving extracellular matrix degradation. NP plays an important role in maintaining the biomechanical properties of the intervertebral discs in the course of IDD [3]. In addition, as the primary composition in the extracellular matrix, proteoglycans, aggrecan and collagens play a vital role in maintaining the normal functions of the intervertebral discs. In fact, the loss of aggrecan and collagen functions has been correlated with the onset of IDD [4]. Furthermore, the balance between MMPs and TIMPs is possibly involved in the pathogenesis and progression of IDD [5]. For example, Deng et al. found that the expression level of MMP-1 in degenerated discs is significantly increased [6]. In addition, Baillet et al. found that an increased level of MMP-1 expression could be induced by exposing the intervertebral discs to surgical injuries [7]. Xu et al. found that the level of MMP-1 expression is significantly correlated with older age as well as the severity of IDD [8]. Overall, the above results suggested that MMP-1 plays an important role in the onset of IDD.
Past studies demonstrated that short micro-RNAs, also termed miRNAs, are implicated functionally in the control of many biological and physiological activities, including cell differentiation, cell apoptosis, and cell proliferation [9]. In addition, miRNAs are also implicated in many human disorders, such as the onset of cardiovascular diseases as well as tumorigenesis [10]. By binding to the 3′ untranslated region (3′ UTR) of their target genes, miRNAs reduce the expression levels of these genes [11–13]. Nevertheless, few studies have focused on the regulatory relationship between the pathogenesis of IDD and miRNAs [14]. A miRNA named miR-202-3p is suspected to play a key role in the pathogenesis of IDD. For example, the reduced expression of miR-202-3p in the presence of IL-1β is associated with the increased expression level of MMP-1 in neuronal precursor cells (NPCs). In addition, by inducing the expression of MMP-1, IL-1β can activate MAPK as well as NF-κB signaling to promote the progression of IDD. On the other hand, a reduced expression level of miR-202-3p remarkably increases the expression level of MMP-1.
The H3K27me3 family of demethylases contains several key family members, such as KDM7A, KDM6A, and KDM6B, all of which can interact with the tri-methyl groups in their targets to specifically overcome certain factors leading to gene repression during tissue differentiation. As a compound frequently used as a nutritional supplement, metformin (MET) treatment has been reported to stabilize the level of circulating PCSK9 in stable coronary artery disease (SCAD) patients [5]. Also, MET can directly act on the activation of certain key signaling pathways involved in senescence and aging, including the UTX/KDM6A pathway [15]. In addition, MET can suppress the demethylating function of UTX and KDM6A enzymes.
MiR-202-3p is involved in the pathogenesis of IDD via regulating MMP-1 expression in NPCs [16]. Furthermore, as an activator of miR-202-3p and KDM6A, MET may regulate the expression of SOX9 via regulating the methylation status of the SOX9 promoter [15, 17]. In this study, we set up an animal model of IDD and treated it with MET to probe the effect of MET on the signaling pathway of KDM6A/SOX9/miR-202-3p/MMP-1 and the treatment of IDD.
Material and methods
Establishment of an animal model of IDD via the noninvasive induction of cumulative axial load
A total of 36 New Zealand male white rabbits (about 5 months of age, weighing 2.2–2.6 kg) were acquired from our laboratory animal center and housed in our animal facility for 7 days of adaptation. Afterwards, the rabbits were assigned randomly to three treatment groups, i.e., 1 – an NC group; 2 – an IDD group; and 3 – an IDD + MET group, with 12 rabbits in each group. During the experiment, all rabbits used to establish the IDD model (i.e., groups 2 and 3) were housed in plastic cages and their posture was maintained in an upright position, and a collar load with a weight of 600 g was hanging on the neck of each rabbit to provide a load induction, which was carried out for 8 h every day. After 8 weeks of induction, the successful establishment of a rabbit IDD model was confirmed by the procedures described below. In the control group, the rabbits were not treated by load induction. In the IDD + MET group, the IDD rabbits were further treated for these 8 weeks with a dose of 30 mg/kg/day of MET given orally. During the entire experiment, the temperature in the animal facility was maintained at 22 ±3°C, while the humidity in the animal facility was maintained at 55 ±5%. The entire experiment was carried out in compliance with the Guideline for the Care and Use of Laboratory Animals.
Culture of neuronal precursor cells
Neuronal precursor cells were isolated from NP tissue samples using a method previously described [18]. In brief, all tissue samples were rinsed with PBS, then digested at 37°C for 30 min using F12 medium (Gibco, Grand Island, NY) containing 2 U/ml of protease. After being treated at 37°C for 4 h using 0.25 mg/ml of type II collagenase (Gibco, Grand Island, NY), the lysate of NP tissue samples was prepared into single cell suspensions and then transferred to a 50 µm cell strainer (BD Biosciences, Franklin Lakes, NJ) before the lysate was centrifuged for 10 min at 1000 g. Finally, the isolated NPCs were suspended in a DMEM medium with 1% L-glutamine, 100 µg/ml streptomycin, 100 U/ml penicillin, as well as 10% FBS (Gibco, Grand Island, NY), and cultured for 1 week under 5% CO2 and 37°C before they were collected for subsequent analyses.
Stimulation of neuronal precursor cells
To stimulate NPCs, they were starved for 12 h in a medium containing no FBS and then exposed to 6 h of IL-1β (5 ng/ml, Peprotech, Rocky Hill, NJ) treatment to stimulate the cells [19].
Bisulfite sequencing
Bisulfite real-time PCR sequencing was carried out to determine the level of DNA methylation in the promoter of the SOX9 gene in collected tissue and cell samples. In brief, the samples were first immersed in a buffer containing 0.1 mM of EDTA and 10 mM of Tris, and then treated with a Bioruptor sonicator to fragment the DNA in the samples. In the next step, the fragmented DNA was precipitated using ethanol, treated using an End-It DNA Repair assay kit (Illumina, Hayward, CA) in accordance with the instructions of the manufacturer to repair the end of the DNA sample, and re-purified using a MinElute DNA clean up assay kit (Qiagen, Hilden Germany) in accordance with the instructions of the manufacturer before carrying out the ligation reaction using a Quick Ligation assay kit (NEB, Ipswich, MA) in accordance with the instructions of the manufacturer. The ligated DNA was then separated using a 5% agarose gel (Bio-Rad, Hercules, CA) and then treated using a MethylEasy assay kit (Bio-Rad, Hercules, CA) in accordance with the instructions of the manufacturer. Finally, the DNA methylation status in the promoter of the SOX9 gene was determined via real-time PCR using a Platinum Taq buffer (Invitrogen, Carlsbad, CA) containing 1 U of Platinum Taq reagent, 2.5 mM of MgCl2 and 50 µM of dNTPs. The PCR conditions were set up in accordance with the instructions of the kit manufacturer.
RNA isolation and real-time PCR
Real-time PCR was carried out to measure the mRNA expression of KDM6A, SOX9, MMP-1, and miR-202-3p in various samples. In brief, total RNA content in each sample was extracted using a Trizol reagent (Invitrogen, Carlsbad, CA) in accordance with the instructions of the kit manufacturer. Then, extracted total RNA was converted to cDNA by reverse transcription, and the mRNA expression of KDM6A (forward primer: 5′-AGCGCAAAGGAGCCGTGGAAAA-3′; reverse primer: 5′-GTCGTTCACCATTAGGACCTGC-3′), SOX9 (forward primer: 5′-AGGAAGCTCGCGGACCAGTAC-3′; reverse primer: 5′-GGTGGTCCTTCTTGTGCTGCAC-3′), MMP-1 (forward primer: 5′-ATGAAGCAGCCCAGATGTGGAG-3′; reverse primer: 5′-TGGTCCACATCTGCTCTTGGCA-3′), and miR-202-3p (forward primer: 5′-TTCCTATGCATATACTTCTT-3′; reverse primer: 5′-GAACATGTCTGCGTATCTC-3′) in each sample was measured using TaqMan real-time PCR (Applied Biosystems, Foster City, CA).
Cell culture, treatment and transfection
The NPCs culture was carried out under 5% CO2, saturated humidity and 37°C. When the cells were confluent, they were randomly divided into 3 groups, i.e., 1. untreated group; 2. IL-1β group (5 ng/ml); and 3. IL-1β (5 ng/ml) + MET group (1 µM). The cells in various groups were treated with designated concentrations of IL-1β and MET or PBS for 12 h before they were collected for subsequent assays. In another set of experiments, NPCs were randomly divided into another 3 groups, i.e., 1 – a scramble control group; 2 – a miR-202-3p precursor group; and 3 – an MMP-1 siRNA group. The cells in various groups were treated with different nucleotides for 48 h using Lipofectamine 3000 (Invitrogen, Carlsbad, CA) in accordance with the instructions of the kit manufacturer before the cells were collected and the expression of target genes in the cells was measured.
Vector construction, mutagenesis and luciferase assay
The results of our previous sequence analysis indicated that miR-202-3p could bind to the 3′ UTR of MMP-1. In order to further explore the regulatory role of miR-202-3p in MMP-1 expression, a pcDNA luciferase plasmid (Promega, Madison, WI) containing the wild type 3′ UTR of MMP-1 was established. At the same time, site-directed mutagenesis was carried out in the miR-202-3p binding site located in the 3′ UTR of MMP-1 using a Quick Change site directed mutagenesis assay kit (Stratagene, San Diego, CA) in accordance with the instructions of the kit manufacturer to generate the mutant type sequence of the 3′ UTR of MMP-1, which was also inserted into a pcDNA luciferase plasmid to establish the luciferase plasmid containing the mutant type 3′ UTR of MMP-1. In the next step, both the mutant type and wild type 3′ UTR of MMP-1 were co-transfected into NPCs in conjunction with either a miR-202-3p mimic or a miR-202-3p inhibitor. The luciferase activity of transfected cells was measured 48 h later using a Bright Glo luciferase assay kit (Promega, Madison, WI) in accordance with the instructions of the kit manufacturer.
Western blot analysis
Western blot was carried out to measure the protein expression of KDM6A, SOX9, and MMP-1. In brief, tissue and cell samples were exposed to primary anti-KDM6A, anti-SOX9, and anti-MMP-1 antibodies (1 : 100 dilution, Abcam, Cambridge, MA) at 4 degrees overnight and then incubated for 2 h at 37°C with horseradish peroxidase conjugated secondary IgG antibodies (Cell Signaling Technology, Danvers, MA) before the samples were visualized using a Clarity Western ECL Substrate (Bio-Rad, Hercules, CA) to determine the protein expression of KDM6A, SOX9, and MMP-1 in each sample.
Statistical analysis
All experimental data were analyzed using Version 5 Expert StatView statistical software (SAS, Cary, NC). The data were analyzed by t-test and one-way analysis of variance (ANOVA). The results were presented as mean ± standard deviation. The level of significance was set to 0.05.
Results
MET treatment alleviated deregulation of KDM6A, SOX9, MMP-1 and miR-202-3p in NPCs
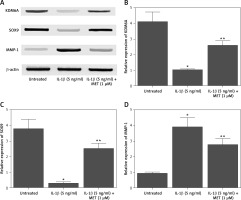
NPCs were treated with IL-1β alone or in combination with MET before Western blot was performed to analyze the differential expression of KDM6A, SOX9 and MMP-1. IL-1β treatment remarkably repressed the expression of KDM6A (Figures 1 A, B) and SOX9 (Figures 1 A, C), while the treatment with MET restored the normal expression of KDM6A and SOX9 to a certain level. In contrast, the expression of MMP-1 was dramatically enhanced by IL-1β while the MET treatment reduced the expression of MMP-1 in NPCs exposed to IL-1β (Figures 1 A, D).
Figure 1
IL-1β induced deregulation of KDM6A, SOX9 and MMP-1 protein expression, and MET treatment reversed the effect of IL-1β (*p < 0.05, vs. untreated group; **p < 0.05, vs. IL-1β group). A – Western blot analysis showed that IL-1β inhibited the expression of KDM6A and SOX9 while activating the expression of MMP-1 protein in NPCs, and MET treatment reversed the effect of IL-1β. B – Quantitative analysis showed that IL-1β inhibited expression of KDM6A protein in NPCs and MET treatment reversed the effect of IL-1β. C – Quantitative analysis showed that IL-1β inhibited expression of SOX9 protein in NPCs and MET treatment reversed the effect of IL-1β. D – Quantitative analysis showed that IL-1β activated expression of MMP-1 protein in NPCs and MET treatment reversed the effect of IL-1β
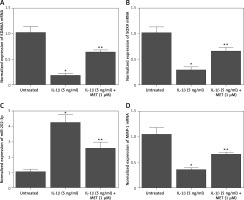
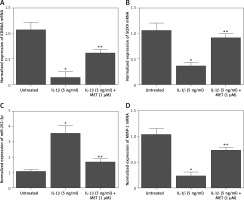
Additionally, quantitative real-time PCR was performed to evaluate the mRNA expression of KDM6A, SOX9, MMP-1, and miR-202-3p in NPCs placed under different conditions. The expression of KDM6A (Figure 2 A), SOX9 (Figure 2 B) and MMP-1 (Figure 2 D) mRNA was notably repressed by IL-1β treatment, while the treatment with MET effectively restored the normal expression of KDM6A, SOX9 and MMP-1 mRNA in IL-1β-treated NPCs. However, the expression of miR-202-3p was apparently elevated by IL-1β treatment, while the treatment with MET reduced miR-202-3p expression in NPCs exposed to IL-1β (Figure 2 C).
Figure 2
IL-1β induced deregulation of KDM6A, SOX9, miR-202-3p and MMP-1 mRNA expression while MET treatment reversed the effect of IL-1β (*p < 0.05, vs. untreated group; **p < 0.05, vs. IL-1β group). A – IL-1β repressed KDM6A mRNA expression in NPCs and MET treatment reversed the effect of IL-1β. B – IL-1β repressed SOX9 mRNA expression in NPCs and MET treatment reversed the effect of IL-1β. C – IL-1β activated miR-202-3p expression in NPCs and MET treatment reversed the effect of IL-1β. D – IL-1β repressed MMP-1 mRNA expression in NPCs and MET treatment reversed the effect of IL-1β
MET treatment reduced hypermethylation of the SOX9 promoter in NPCs
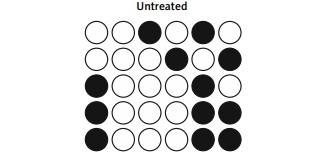
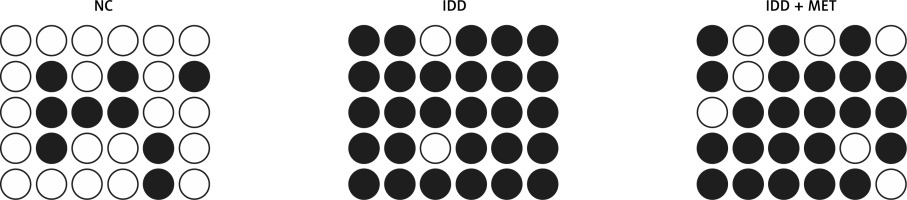
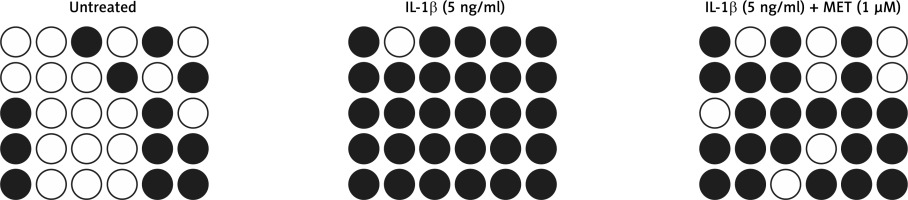
DNA methylation plays an important role in gene regulation. In this study, we performed bisulfite sequencing PCR to analyze the different levels of DNA methylation of the SOX9 promoter in NPCs exposed to different conditions. IL-1β treatment significantly increased DNA methylation of the SOX9 promoter and down-regulated SOX9 expression. MET treatment notably suppressed hypermethylation of the SOX9 promoter in NPCs (Figure 3).
MiR-202-3p repressed expression of MMP-1 through binding to its 3′ UTR
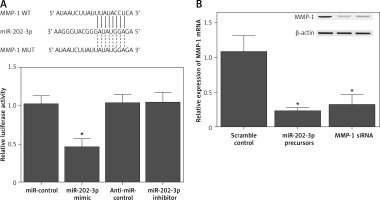
Sequence analysis indicated that miR-202-3p could bind to the 3′ UTR of MMP-1. In order to further explore the regulatory role of miR-202-3p in MMP-1 expression, a luciferase plasmid containing the MMP-1 3′ UTR was established and co-transfected into NPCs in conjunction with either a miR-202-3p mimic or a miR-202-3p inhibitor. The luciferase activity of MMP-1 was significantly repressed by the miR-202-3p mimic and enhanced by the miR-202-3p inhibitor (Figure 4 A). Quantitative real-time PCR and Western blot were performed to evaluate the expression of MMP-1 mRNA and protein in NPCs transfected with a miR-202-3p precursor and MMP-1 siRNA, respectively. The expression of MMP-1 mRNA and protein was notably inhibited by both the miR-202-3p precursor and MMP-1 siRNA (Figure 4 B).
Figure 4
MiR-202-3p inhibited the expression of MMP-1 through binding to its 3′ UTR. A – The luciferase activity of MMP-1 vector was inhibited by miR-202-3p mimics and increased by miR-202-3p inhibitors (*p < 0.05, vs. miR-controls). B – The expression of MMP-1 protein was effectively inhibited by miR-202-3p precursors and MMP-1 siRNA (*p < 0.05, vs. scramble controls)
MET administration restored the abnormal expression of KDM6, SOX9, MMP-1 and miR-202-3p in IDD rabbits
An IDD (intervertebral disc degeneration) rabbit model was established and the value of disc height index (DHI) was measured at different time points. In the control rabbits, the DHI was progressively decreased. However, the DHI for IDD rabbits remained stable at different time points, indicating the abnormal elevation of DHI when compared with the control. MET administration could restore the normal value of DHI in IDD rabbits to a certain degree (Figure 5 A). Additionally, we performed Western blot to evaluate the expression of KDM6, SOX9 and MMP-1 protein in IDD rabbits. The abnormally up-regulated KDM6 (Figure 5 B) and SOX9 (Figure 5 C) expression as well as the abnormally down-regulated MMP-1 (Figure 5 D) expression was restored to normal by MET treatment. Moreover, the qPCR results showed that MET treatment decreased the expression of KDM6A (Figure 6 A), SOX9 (Figure 6 B) and MMP-1 (Figure 6 D) mRNA, while increasing the expression of miR-202-3p (Figure 6 C) in IDD rabbits. Similar to the change of SOX9 expression in IDD rabbits, the DNA methylation of the SOX9 promoter was significantly increased in IDD rabbits, while MET administration efficiently reduced the hypermethylation of the SOX9 promoter in IDD rabbits (Figure 7).
Figure 5
MET treatment effectively restored the deregulation of KDM6A, SOX9 and MMP-1 protein expression in IDD rabbits (*p < 0.05, vs. untreated group; **p < 0.05, vs. IL-1β group). A – Graphic representation of DHI of rabbit discs. B – Western blot analysis showed that the inhibited expression of KDM6A and SOX9, and activated expression of MMP-1 protein, in IDD rabbits were recovered by MET treatment. C – Quantitative analysis showed that the inhibited expression of KDM6A protein in IDD rabbits was recovered by MET treatment. D – Quantitative analysis showed that the inhibited expression of SOX9 protein in IDD rabbits was recovered by MET treatment. E – Quantitative analysis showed that the activated expression of MMP-1 protein in IDD rabbits was recovered by MET treatment
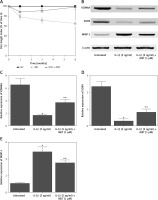

Figure 6
The deregulated mRNA expression of KDM6A, SOX9, MMP-1 and miR-202-3p in IDD rabbits was recovered by MET treatment (*p < 0.05, vs. untreated group; **p < 0.05, vs. IL-1β group). A – Repressed expression of KDM6A mRNA in IDD rabbits was restored by MET treatment. B – Repressed expression of SOX9 mRNA in IDD rabbits was restored by MET treatment. C – Enhanced expression of miR-202-3p in IDD rabbits was restored by MET treatment. D – Repressed expression of MMP-1 mRNA in IDD rabbits was restored by MET treatment
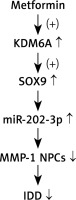
In conclusion, our results revealed that MET treatment up-regulated KDM6A expression, which then promoted the demethylation of the SOX9 promoter to up-regulate its expression and induce the expression of miR-202-3p. Increased expression of miR-202-3p inhibited the expression of MMP-1 and attenuated the severity of IDD (Figure 8).
Discussion
As a process involving both senescence and apoptosis, IDD can be alleviated by MET via stimulating the activity of autophagy. MET may exert its effects on inhibiting the demethylating activity of H3 Lys27 enzymes by interacting with the protein residues residing in the catalytic domain of UTX and KDM6A. In this study, we treated NPCs with IL-1β to establish a cellular model of IDD. We also constructed an IDD rabbit model to explore the therapeutic efficiency of MET in IDD treatment. Furthermore, we carried out bisulfite sequencing PCR to evaluate the level of DNA methylation of the SOX9 promoter in the IDD models. MET treatment could effectively reduce the level of SOX9 promoter methylation in NPCs and IDD rabbits stimulated by IL-1β.
KDM6A, also termed lysine specific demethylase 6A and UTX, acts as a demethylase specific to histone H3K27. In fact, KDM6A plays a critical role in regulating the differentiation of stem cells toward M2 macrophages [20]. In addition, the over-expression of KDM6A in MSCs can promote osteogenesis while inhibiting adipogenesis [21]. In fact, the methylation of histone lysine is under the control of histone lysine demethylase (KDM) and histone lysine methyltransferase (KMT).
SOX9 is known as an important transcription factor involved in chondrogenic differentiation. Past results demonstrated that the level of SOX9 protein expression is apparently reduced after the expression of KDM6A is knocked down in PDLSC cells. In fact, as a key regulator involved in the differentiation of Sertoli cells, SOX9 can affect the expression of a wide range of target genes in both human and mouse [22–25]. In addition, SOX9 was shown previously to control the expression level of pri-miR-140 involved in activation of the Wwp2 gene [26, 27].
Growing evidence suggests that a wide range of miRNAs are involved in the onset of IDD by regulating the extent of nucleus pulposus cell apoptosis as well as degradation of the extracellular matrix. In addition, the expression level of miR-30d is negatively associated with the expression level of SOX9 in NP tissues. On the other hand, the expression level of miR-202-5p/3p is low in mice with the Sox9 gene knocked out, indicating that pri-miR-202 acts as a downstream target of SOX9. In this study, we performed luciferase assays to explore the inhibitory role of miR-202-3p in MMP-1 expression. The luciferase activity of the MMP-1 vector was significantly repressed by miR-202-3p mimics and increased by miR-202-3p inhibitors. As a regulator in the expression of MMP-1, miR-202-3p expression can suppress the expression level of MMP-1 in the presence of IL-1β. On the other hand, transfection using miR-202-3p inhibitors apparently promoted the production of MMP-1, suggesting that miR-202-3p can reduce the expression level of MMP-1 in NP cells to participate in the onset of IDD.
As enzymes participating in the extracellular matrix degradation, MMPs, especially MMP-1, can cleave endogenous type I, type II, as well as type III collagens in the interstitial space, leading to possible disc degeneration and alterations in the disc structure [18, 28–30]. In addition, the expression levels of MMP-3 and MMP-1 are both increased during the onset of lumbar disc degeneration [31]. In a similar study, it was found that the level of MMP-1 expression was significantly associated with age [8].
In conclusion, the findings of this study demonstrated that miR-202-3p is involved in the pathogenesis of IDD via regulating MMP-1 expression in NPCs. Furthermore, as an activator of miR-202-3p and KDM6A, MET may regulate the expression of SOX9 via regulating the methylation status of the SOX9 promoter. Meanwhile, treatment with MET may enhance KDM6A expression. In this study, we set up an animal model of IDD and treated it with MET to probe the effect of MET on the signaling pathway of KDM6A/SOX9/miR-202-3p/MMP-1 and the treatment of IDD.